For a complete publication list, please refer HERE or to my Google Scholar profile.
A selected collection of my (co)first-authored publications is shown below.
6. Integrative multi-omics deciphers the spatial characteristics of host-gut microbiota interactions in Crohn’s disease [Cell Reports Medicine] (Co-first, 20230620)
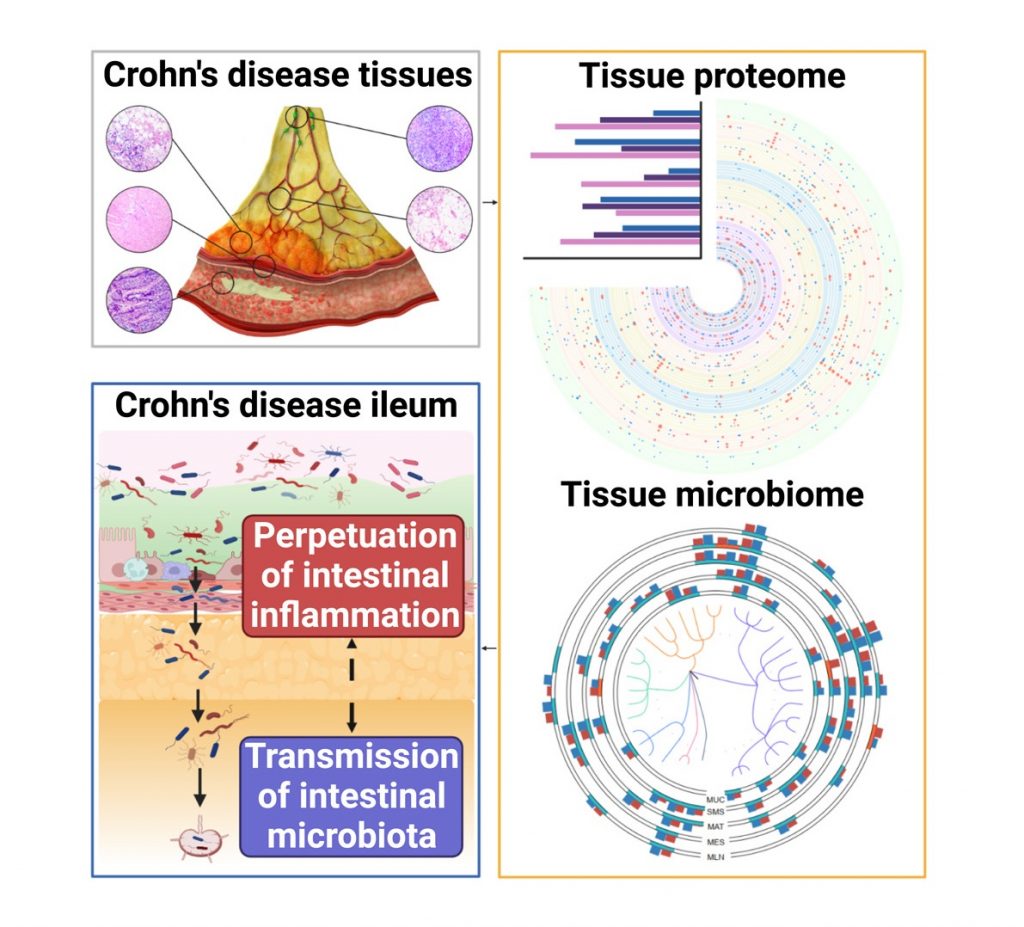
Host-microbial interactions play critical roles during gut inflammation in Crohn’s Disease (CD), a type of Inflammatory Bowel Disease (IBD).
We applied proteomics and microbiomics to characterize five sections of diseased tissue from thirty CD patients, and thereby analyzed the host responses as well as their interactions with bacterium. Over 600 proteomes were analyzed in the project and we specially implemented Pulse DIA technique on a TimsTOF mass spectrometry to increase proteome depth. We proposed diagnostic biomarkers with signature alteration that are also shown in serum and feces.
I contributed to proteomic experiments, data analysis, and paper revision.
克罗恩病(CD)是一种复杂的炎症性肠病,其发病过程中,宿主与微生物之间的相互作用十分关键但其机制尚未被充分表征。
我们对 30 名 CD 患者回肠中的五处病变组织切片进行了蛋白质组和微生物组的分析。其中,蛋白质组数据基于新近开发的pulse-diaPASEF方法采集、采集总数目超过 600 个。从超过8000种蛋白结果的挖掘中,我们提炼出了一批与CD特征显著相关的诊断型生物标志物,并通过额外探索发现这些标志物也可从血清和粪便中被表征和鉴定。
我负责了蛋白质组学实验、数据分析和论文修订的工作。
5. Proteomics Investigation of Diverse Serological Patterns in COVID-19 [Molecular & Cellular Proteomics] (First, 20230105)

Our body relies heavily on the secretion of serum antibodies to defend against SARS-CoV-2 attacks. From current clinical reports, we noticed patients with unexpected serological patterns, such as negative antibody expression throughout COVID-19, and exceptionally high expression of IgM or IgG at their plateau. These observations suggest diverse host responses during COVID-19, which have yet to be assessed.
In this study, we applied two-year clinical manifestation and lop、ngitudinal serum proteomics to understand the serology in a cohort of 144 COVID-19 patients. The findings suggest that COVID-19 patients who do not express antibodies developed cellular immunity for viral defense, and that high titers of IgM might not be favorable to COVID-19 recovery.
I contributed to proteomic experiments, data analyses, and manuscript writing.
人体主要依赖血清抗体的分泌来抵御新冠病毒侵袭。但现有临床报告显示,部分患者呈现特殊的血清学特征:有的在整个病程中始终未检出抗体表达,有的则出现异常高水平的IgM或IgG平台期。这些现象提示新冠病毒感染过程中存在多样化的宿主免疫应答机制,其临床意义尚待系统评估。
本研究通过整合144名新冠患者为期两年的临床表现与纵向血清蛋白质组数据,首次揭示:未产生抗体的患者可能通过细胞免疫实现病毒防御,而高滴度IgM反而可能不利于临床康复。
我负责了蛋白质组学实验、数据解析及论文撰写工作。
Affiliated blog相关博客
4. A Common Mechanism of Temperature-sensing in ThermoTRP Channels [Under review] (Co-first, 20220524)
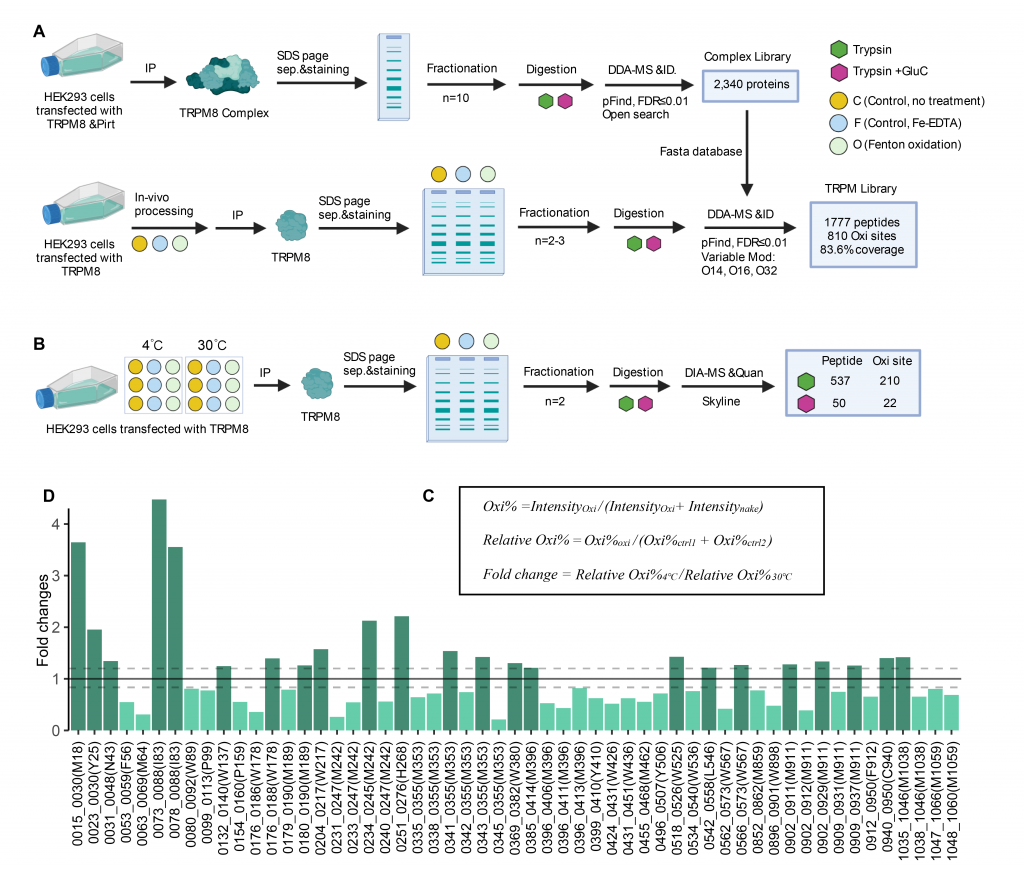
Detecting temperature is crucial for the survival of living organisms. Thermo transient receptor potential (thermoTRP) channels, such as TRPV1 or TRPM8, have been identified as prototypic heat or cold sensors, respectively. However, how they detect temperature remains elusive.
We developed hydroxyl radical footprinting-data-independent acquisition mass spectroscopy (HRF-DIA-MS), a combinative technology, to quantitatively profile the protein residues that undergo burial/exposure conformational rearrangements during temperature activation. Our findings established that the water-protein interaction is a common mechanism underlying temperature sensing in TRPM8 and TRPV1.
I contributed to the proteomic methodology and data analysis.
3. Pulmonary and Renal Long COVID at Two-year Revisit [iScience] (Co-first, 20240620)
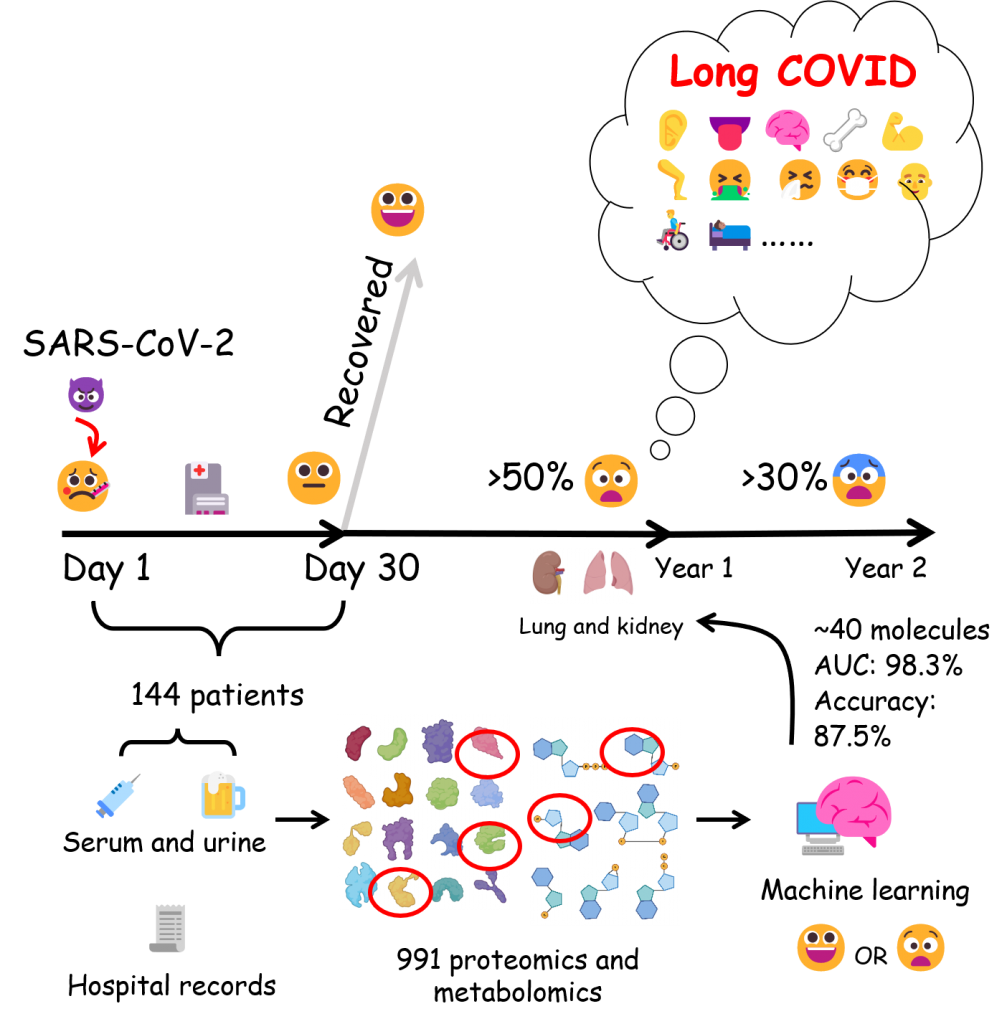

More than 450 million individuals have recovered from COVID-19, but little is known about the host responses to long COVID.
We integrated a multi-omics resource containing proteomic and metabolomic analyses of 991 blood and urine specimens from 144 COVID-19 patients. Our data depicts the longitudinal clinical and molecular landscape of COVID-19 with up to two-year follow-up and presents a method to predict pulmonary and renal long COVID.
I contributed to multi-omics experiments, data analyses, and manuscript writing.
3-min video illustration
2. Computational Optimization of Spectral Library Size Improves DIA-MS Proteome Coverage and Applications to 15 Tumors [Journal of Proteome Research] (Co-first, 20211108)

Public MS data repositories are being generated for proteomic studies. They provide accessible spectral information for library-based DIA-MS data analyses. However, vast number of false positives may be introduced due to an enlarged search space, leading to an unpleasant proteomic identification.
Via a two-step strategy called subLib to generate experiment-specific subset libraries from the public data, we downscaled the library size while enhanced the resultant proteomic coverages from DIA-MS data by 30-40%. This method is proven handy in proteomics of 15 tumor tissues.
I contributed to data revision and manuscript writing.
1. Proteomic and Metabolomic Investigation of Serum Lactate Dehydrogenase Elevation in COVID-19 Patients [Proteomics] (Co-first, 20210513)
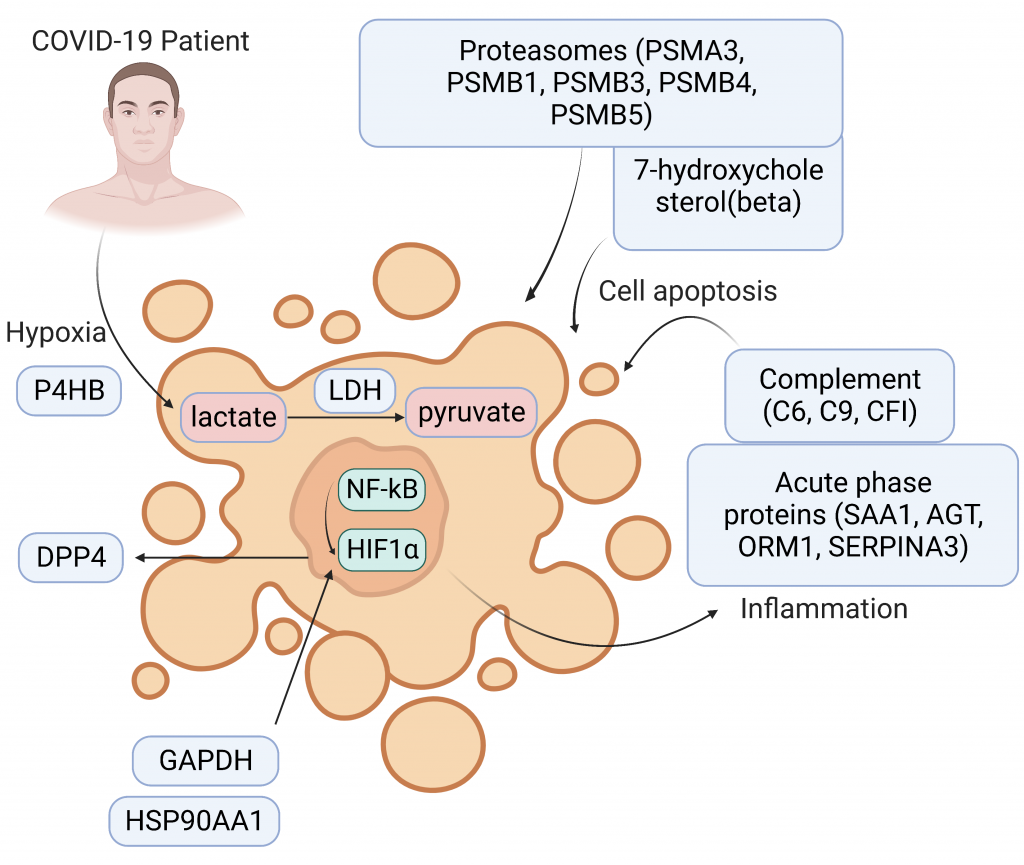
Top Cited Article 2021-2022 in Proteomics
With many risk factors proposed for COVID-19, biological understandings of their mechanisms are in lack. We extracted the published proteomic and metabolomic data to investigate the abnormal elevation of lactate dehydrogenase (LDH), a recognized molecular risk factor in COVID-19 patients. Not only is LDH expression levels associated with COVID-19 severity, its dysregulation associates with organ injuries and hypoxia responses.
The ideas and methods applied in this study could be of reference to other risk factor studies for a more credible medical determinization.
I contributed to data analyses, interpretation, and manuscript writing.
Patents
- 郭天南;梁潇;朱怡. 伴随蛋白/肽在提高蛋白质组实验效率中的应用. ZL202010762585.6 (CN patent) certificate
- 郭天南;梁潇;石颖秋;王瑛睿. 一种高效清除液相色谱中多肽残留的方法及应用. ZL202210103388.2 (CN patent) certificate
- 郭天南;梁潇;陈晨;石颖秋. 一种生物样本的蛋白质组学分析方法. ZL202210399034.7 (CN patent) certificate
- 郭天南;梁潇. 一种分析细胞亚型的方法. 审核中 (CN patent pending; Pub. No. CN117309727A)